Laboratory Development Test Validation of an H&E-Based Deep Learning Histogenomic Model to Predict Biomarkers in Endometrial Cancer
USCAP 2026
Qiyuan Hu, Kshitij Ingale, Sun Hae Hong, Rohan Joshi, Jacob Gordon, Yoni Muller, Ben Terdich, Jason Blue-Smith, Ryan Jones, Nike Beaubier, Chithra Sangli, Riccardo Miotto
Background
Next-generation sequencing (NGS) can provide critical information for selecting appropriate therapies for cancer patients and improving outcomes. However, NGS is not always performed, especially when not indicated in clinical guidelines. We developed a digital pathology model to predict biomarkers associated with molecular subtypes of endometrial cancer, including MSI-High status and pathogenic or likely pathogenic (P/LP) alterations in CTNNB1, POLE, PTEN, and TP53 from digital hematoxylin and eosin (H&E) whole slide images (WSIs) and validated the model using a CAP/CLIA laboratory-developed test (LDT).
Design
The model was trained and tuned using 4,917 WSIs (4,809 patients) and validated on 388 WSIs (388 patients) of endometrial cancer. Samples selected were biopsies or surgical resections from any tissue site and diagnosis subtype. The validation cohort was enriched for rare biomarkers, e.g. POLE. Samples were sequenced using the Tempus xT DNA-seq assay, which established the ground truth. We extracted tile embeddings from WSIs using a foundation model, H-optimus-0, and trained a multi-task attention-based multiple instance learning model to predict 17 relevant biomarkers in endometrial cancer. In the LDT validation, the model was evaluated for 5 biomarkers with predefined acceptance criteria. The analytical accuracy study evaluated the model’s ability to accurately predict the biomarkers. The analytical precision study assessed reproducibility and repeatability using inter/intra-scanner rescans. For analytical sensitivity and specificity, we established a limit of detection (LoD) on the tissue area and the model’s robustness to artifacts, respectively.
Results
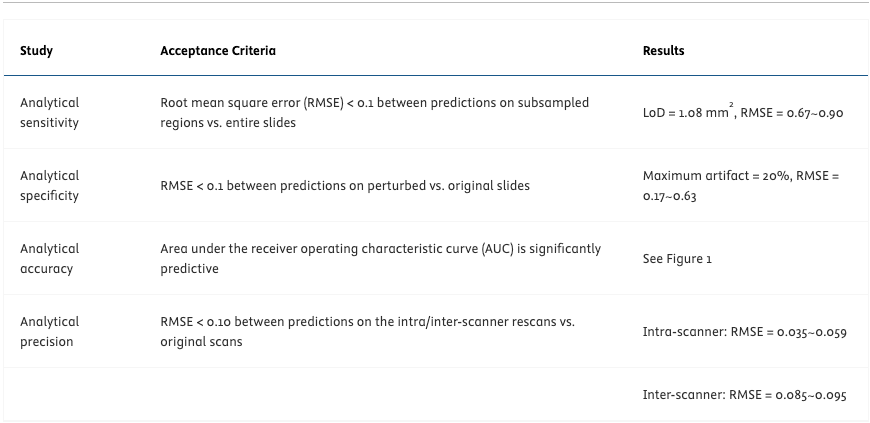
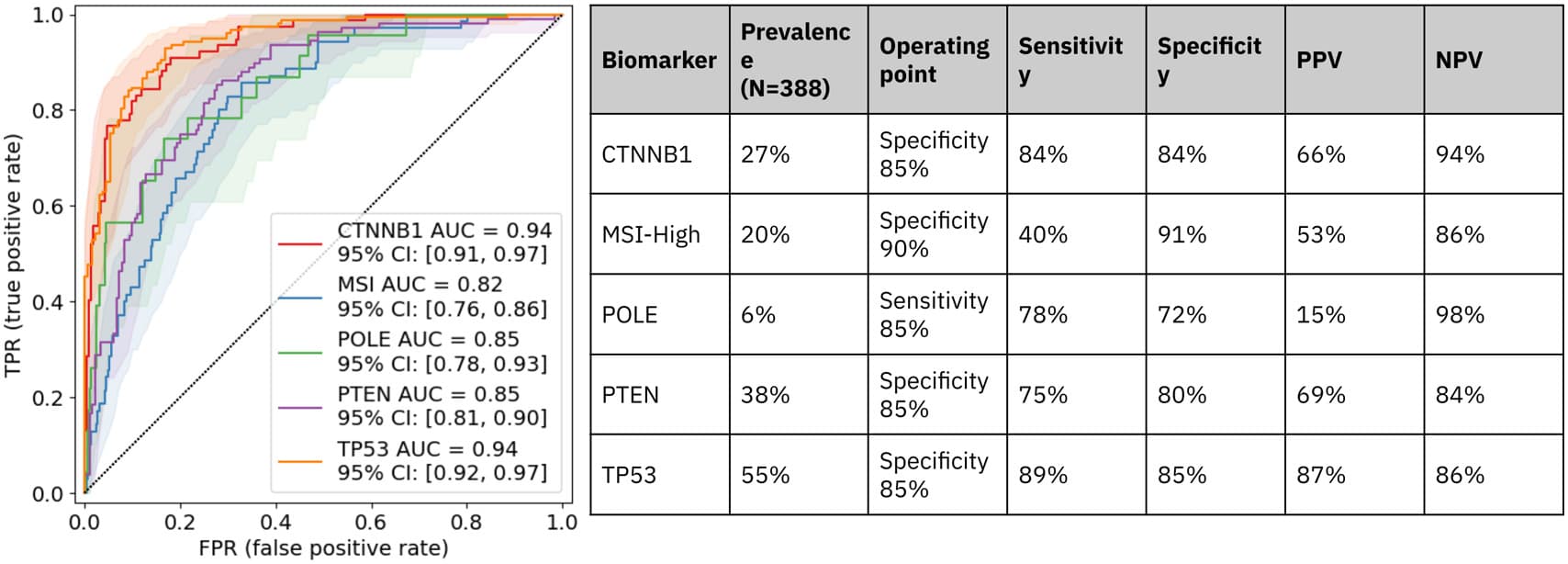
Fig. 1 shows the validation cohort sample count, the model’s AUC, and performance metrics at a predetermined operating point for each biomarker. Table 1 summarizes key results from the LDT validation. The model achieved significantly predictive AUCs for all biomarkers and improved performance compared to previous studies. The discrepancies between model predictions on the original scans vs on inter/intra-scanner rescans satisfied RMSE<0.1 for all biomarkers. The model demonstrated a LoD of 1.08 mm2 tissue and robustness to 20% blur or color artifacts.

Conclusions
This histogenomic model met all acceptance criteria during LDT validation. It has the potential to be deployed for triaging patients for NGS or other confirmatory testing, identifying patients with care gaps, prioritizing tissue, and enriching clinical trial cohorts.
Related publications