-
PROVIDERS
REGISTER NOW
Upcoming Webinar:
MRD data insights across the cancer care continuum -
LIFE SCIENCES
REGISTER NOW
From insight to impact: Leveraging the AI-enabled Next platform with BMS to advance equitable access in precision oncology
-
PATIENTS
It's About Time
View the Tempus vision.
- RESOURCES
-
ABOUT US
View Job Postings
We’re looking for people who can change the world.
- INVESTORS
01/05/2026
A Real-World Study of Claudin 18.2 (CLDN18.2) Association With Molecular Subtypes, Mutations/Biomarkers, Immune Landscapes, and Gene Signatures and Prognostic Value in Pancreatic Ductal Adenocarcinoma (PDAC)
ASCO GI 2026
PRESENTATION
Authors
Guoqiang Zhang, Diarmuid M. Moran, Qunli Xu, Abraham Guerrero
Background:CLDN18.2, a validated therapeutic target and biomarker in gastric cancer, is often highly expressed in PDAC.
Methods:Correlation between CLDN18 protein expression (VENTANA CLDN18 [43-14A] RxDx Assay; Roche Diagnostics) and CLDN18.2 RNA expression (RNAseq assay) was assessed in matched PDAC samples. A real-world clinical genomic cohort of patients with PDAC (Tempus database) was stratified into CLDN18.2 RNA-high, -low, and ‑negative groups by protein–RNA correlation. CLDN18.2 RNA groups were compared for molecular subtypes, mutations, programmed cell death ligand 1 (PD-L1), immune cell proportions, and gene signature scores. Overall survival (OS) of CLDN18.2 RNA groups was compared using Cox proportional hazards regression analyses.
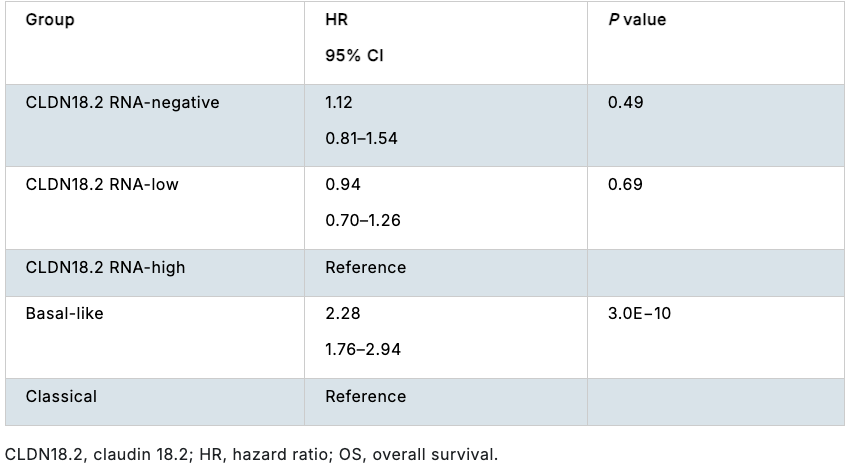
Results:CLDN18 protein and CLDN18.2 RNA showed strong correlation (R=0.89; P=2.2E−16) in 60 matched samples. The threshold for CLDN18.2 RNA-high tumors, corresponding to ≥75% of tumor cells with moderate to strong membranous CLDN18 immunohistochemistry staining, was set at 5.22 log2 transcripts per million (TPM); the threshold for CLDN18.2 RNA-negative tumors was 1 TPM. Baseline demographics were balanced among groups. Basal-like and classical subtypes were more abundant in CLDN18.2 RNA-negative and -high tumors, respectively; CLDN18.2 RNA groups had a moderate to strong association with these molecular subtypes (Cramér’s V=0.48; P=1.6E−43). No notable associations between mutations and CLDN18.2 RNA groups were found in this analysis. A significant but weak negative association was shown between PD-L1 and CLDN18.2 RNA (Cramér’s V=0.22; P=2.0E−7). CLDN18.2 RNA-negative tumors had higher levels of immune cells (B cells, natural killer cells, neutrophils, macrophages, and T cells) as well as immunosuppressive cells (such as regulatory T cells and M2 macrophages) and were associated with higher interferon γ, inflammatory, and mesenchymal gene signatures. Compared with the overall population (N=539), CLDN18.2 RNA-negative patients (n=172) had unfavorable OS and CLDN18.2 RNA-high patients (n=151) had no significant difference in OS. In a multivariable OS analysis (Table), CLDN18.2 was not an independent prognostic factor.
Conclusions:Real-world data suggest that CLDN18.2 is not an independent factor associated with prognosis (OS) in PDAC. Observations regarding the association of CLDN18.2 RNA expression with various molecular and cellular contexts warrant further exploration.
Multivariable OS analysis with CLDN18.2 RNA groups and molecular subtypes as covariates.