-
PROVIDERS
New MRD Medicare Coverage for Select Indications*
*When coverage criteria are met. Additional criteria and exceptions for coverage may apply.
-
LIFE SCIENCES
Register now
UPCOMING WEBINARTranslating data into an actionable R&D strategy
-
PATIENTS
It's About Time
View the Tempus vision.
- RESOURCES
-
ABOUT US
View Job Postings
We’re looking for people who can change the world.
- INVESTORS
03/23/2026
Validation of H&E-based Laboratory Development Test Predicting Molecular Alterations in Non-Small Cell Lung Cancer
USCAP 2026
PRESENTATION
Authors
Kshitij Ingale, Qiyuan Hu, Sun Hae Hong, Rohan Joshi, Jacob Gordon, Yoni Muller, Ben Terdich, Jason Blue-Smith, Ryan Jones, Nike Beaubier, Chithra Sangli, Riccardo Miotto
Background – Molecular alterations detected by next generation sequencing (NGS) are important for identifying patients who could benefit from targeted therapies. However, NGS is not always performed due to cost, tissue availability and long turnaround times. Digital pathology models using routinely generated hematoxylin and eosin (H&E) stained images offer a promising solution to prioritize patients for NGS testing. In this study, we developed and validated an H&E-based model to predict clinically-actionable pathogenic and/or likely pathogenic alterations in ALK, BRAF, EGFR, ERBB2, MET, RET, and ROS1 in non-small cell lung cancer (NSCLC).
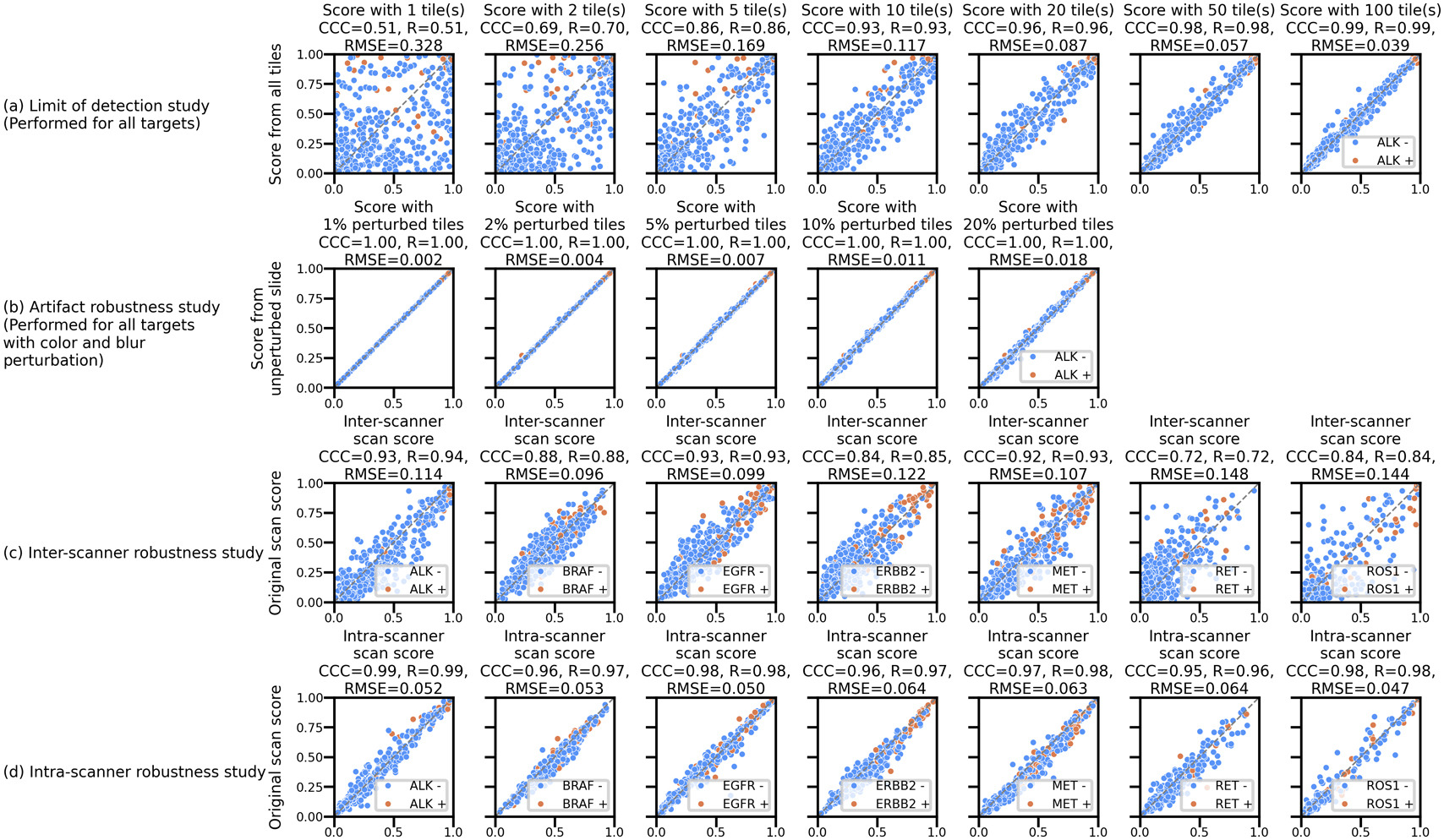
Design – The model was trained using a cohort of 42,467 images (40,990 patients) and later validated on 510 images (510 patients) along with their corresponding labels generated from Tempus xT, an NGS-based DNA assay. A multi-task model was trained to predict the binary labels using an attention based multiple instance learning model with H-optimus-0 foundation model serving as a featurization model. Simulation studies were performed to determine limits for tissue area and artifact content by progressively incorporating more tissue patches and simulated blurry and color perturbed patches during model inference. Model robustness to images obtained from different scanners and multiple scans obtained from the same scanner was also evaluated. Operating point based metrics were calculated on the validation cohort using thresholds determined on a tuning set with similar data characteristics.
Results – The model demonstrated stable performance for cases with tissue area larger than 1.08 mm2 (Figure 1a) and a maximum simulated artifact content of 20% (Figure 1b). Model scores were consistent across different scans from the same scanner (0.047 ≤ Root Mean Square Error (RMSE) ≤ 0.064, Figure 1c), and relatively less consistent across scans from different scanners (0.096 ≤ RMSE ≤ 0.148, Figure 1d). The model significantly predicted alterations (average ROC-AUC = 0.75, Figure 2a) and achieved target sensitivity (and specificity for EGFR) on the validation set using thresholds determined by a tuning set (Figure 2b).
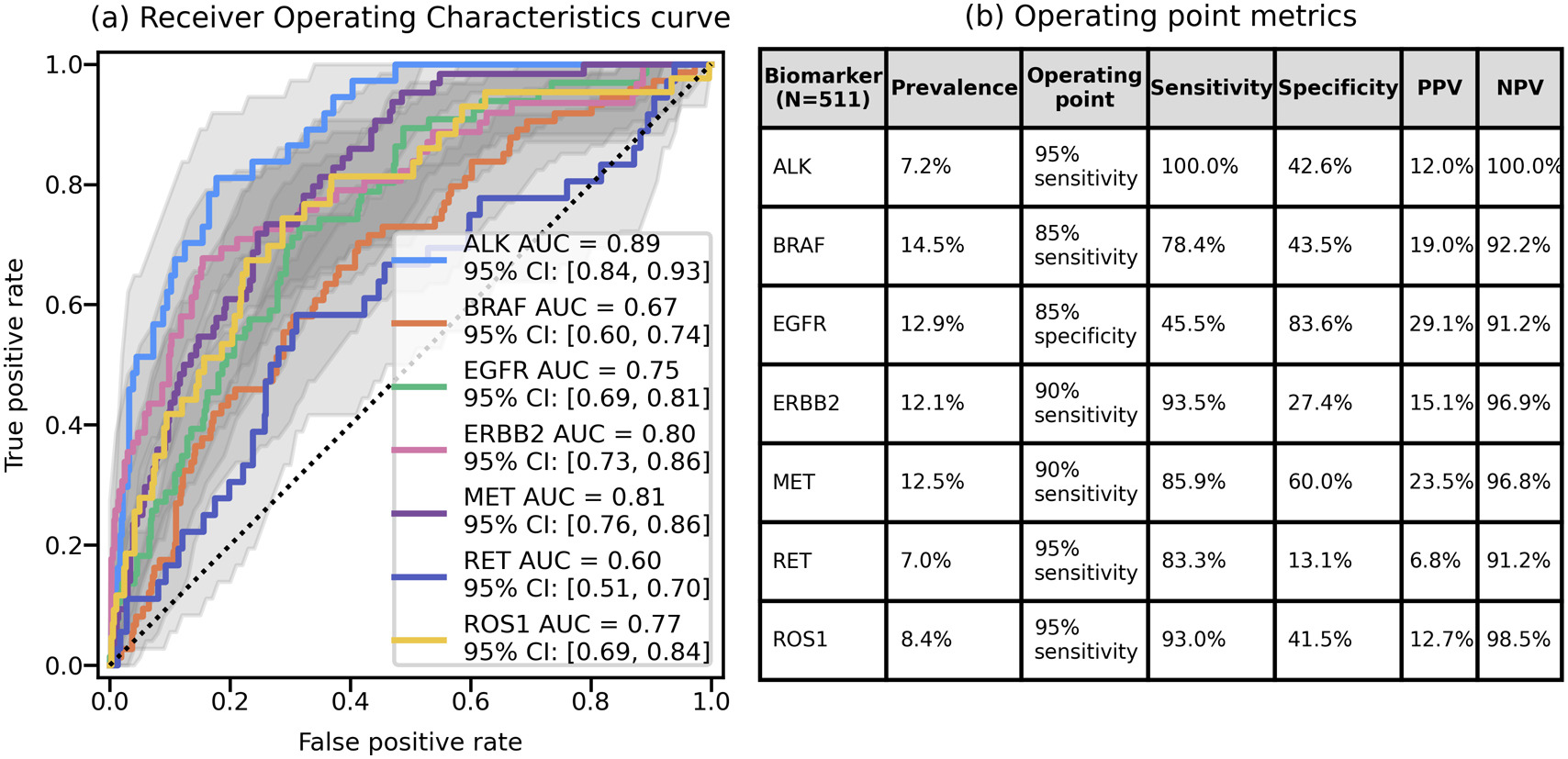
Conclusions – Our study demonstrates that H&E based models can successfully predict actionable molecular alterations. Patients with a high likelihood of these alterations could be alerted and prioritized for confirmatory testing. Further rigorous validation could evaluate their effectiveness in a clinical setting through a prospective trial.