-
PROVIDERS
Learn more
Join Tempus at the 2026 ASCO® Annual Meeting!
-
LIFE SCIENCES
REGISTER NOW
From insight to impact: Leveraging the AI-enabled Next platform with BMS to advance equitable access in precision oncology
-
PATIENTS
It's About Time
View the Tempus vision.
- RESOURCES
-
ABOUT US
View Job Postings
We’re looking for people who can change the world.
- INVESTORS
01/20/2026
From image to insight: How Tempus is leveraging AI to advance biomarker screening at scale
By leveraging Paige Predict to screen for genomic and phenotypic biomarkers, clinicians and researchers have rapid access to valuable information to prioritize testing and pre-screen patients at scale.
Authors
Razik Yousfi
SVP & GM, Imaging, Tempus

Caitlin McWilliams
VP, Strategy & Operations, Tempus

Siqi Liu PhD
VP, Machine Learning, Tempus

SVP & GM, Imaging, Tempus

Caitlin McWilliams
VP, Strategy & Operations, Tempus

Siqi Liu PhD
VP, Machine Learning, Tempus

We have developed Paige Predict for biomarker prediction1, an AI application predicting the status of over 1,600 biomarkers across 505 genes to inform testing prioritization, improve diagnostic yield, and accelerate R&D to identify novel digital biomarkers. At Tempus, we have validated Paige Predict on 123 clinically relevant biomarkers and oncogenic molecular pathways, and it is in use with our xT CDx assay to provide biomarker insights when tissue is insufficient for our solid tumor DNA sequencing workflows.
Cancer is not a single disease but a collection of thousands of distinct molecular conditions. Identifying the specific genomic alterations, or biomarkers, that drive a patient’s tumor is the foundation of precision oncology. Next-generation sequencing (NGS) has revolutionized our ability to profile tumors, while the introduction of novel targeted therapies, clinically actionable biomarkers, and treatment pathways enable increasingly personalized oncology care plans.
However, as a result, medical oncologists and pathologists are faced with an increasing number of diagnostic tests that place significant demand on limited tissue. While AI can identify biomarkers from tissue images2,3,4, the majority of these efforts focus on single biomarkers within a specific tumor type, limiting their generalizability and requiring resource- and time-intensive development effort to expand to a clinically meaningful scale.
This is why Paige, now a Tempus company, built Paige Predict. Paige Predict is a suite of digital pathology AI applications designed for NGS workflows. In this article, we will take a closer look at Paige Predict’s digital biomarker solution that predicts over 1,600 biomarkers across cancer types. Our application analyzes a single, digitized hematoxylin-and-eosin (H&E) stained pathology image, enabling rapid biomarker screening at scale.
Building a high-throughput pan-cancer genetic and phenotyping biomarker panel
Paige Predict is a high-throughput AI application5 capable of detecting a broad panel of genomic biomarkers from H&E images across multiple cancer types. Paige Predict is available for use in common cancers, including tissue-limited indications such as non-small cell lung and pancreatic cancer, and generalizes to rare cancer types and genomic alterations where standalone models are impractical.
This breadth of applicability is enabled by Virchow 2, Paige’s state-of-the-art pathology foundation model. Virchow 2 is pre-trained on 3 million digital pathology slides, capturing diverse morphological patterns across tumor types, tissue sites, and staining conditions. This large-scale pretraining provides a robust representation of histopathologic phenotypes, allowing downstream models to generalize across cancers and institutions.
Paige Predict builds upon Virchow 2 by applying supervised learning on top of its slide-level embeddings using paired histopathology and genomic sequencing data. The primary training cohort includes >40,000 cancer patients from a leading NCI, including pathology images and corresponding molecular results. We also leverage data representing an additional >150,000 patients from Tempus’ vast multimodal database to fine-tune a version of the model, further adapting the predictions to the broad spectrum of clinically relevant targets identified by Tempus’ existing tissue sequencing panels.
To determine performance and generalizability, we evaluated Paige Predict across three separate cohorts, with data derived from >800 institutions and >45 countries, as well as ground truth from multiple different solid tumor sequencing panels. Across datasets, the system consistently demonstrates high performance across a wide range of clinically actionable biomarkers, including:
- Pan-cancer biomarkers, such as microsatellite instability high/deficiency of mismatch repair (MSI-H/dMMR), which are regularly tested for colorectal cancer but not as part of routine screening in numerous other histologies due to low incidence. Detection, including in low-incidence histologies, may indicate vulnerability to certain immuno-oncology drugs.
- Clinically actionable mutations common in certain cancer types, but rare in others. For example, we predict activating BRAF mutations, which have therapeutic significance, across cancer types. These mutations are common in thyroid cancers and melanoma, but are less common in colorectal and non-small cell lung cancers.
- Predictive and prognostic biomarkers in rare cancer types, such as gliomas. In cancers of the central nervous system, Paige Predict shows admirable performance in predicting IDH1 mutations, important for prognosis and therapy selection.
Paige Predict workflow:
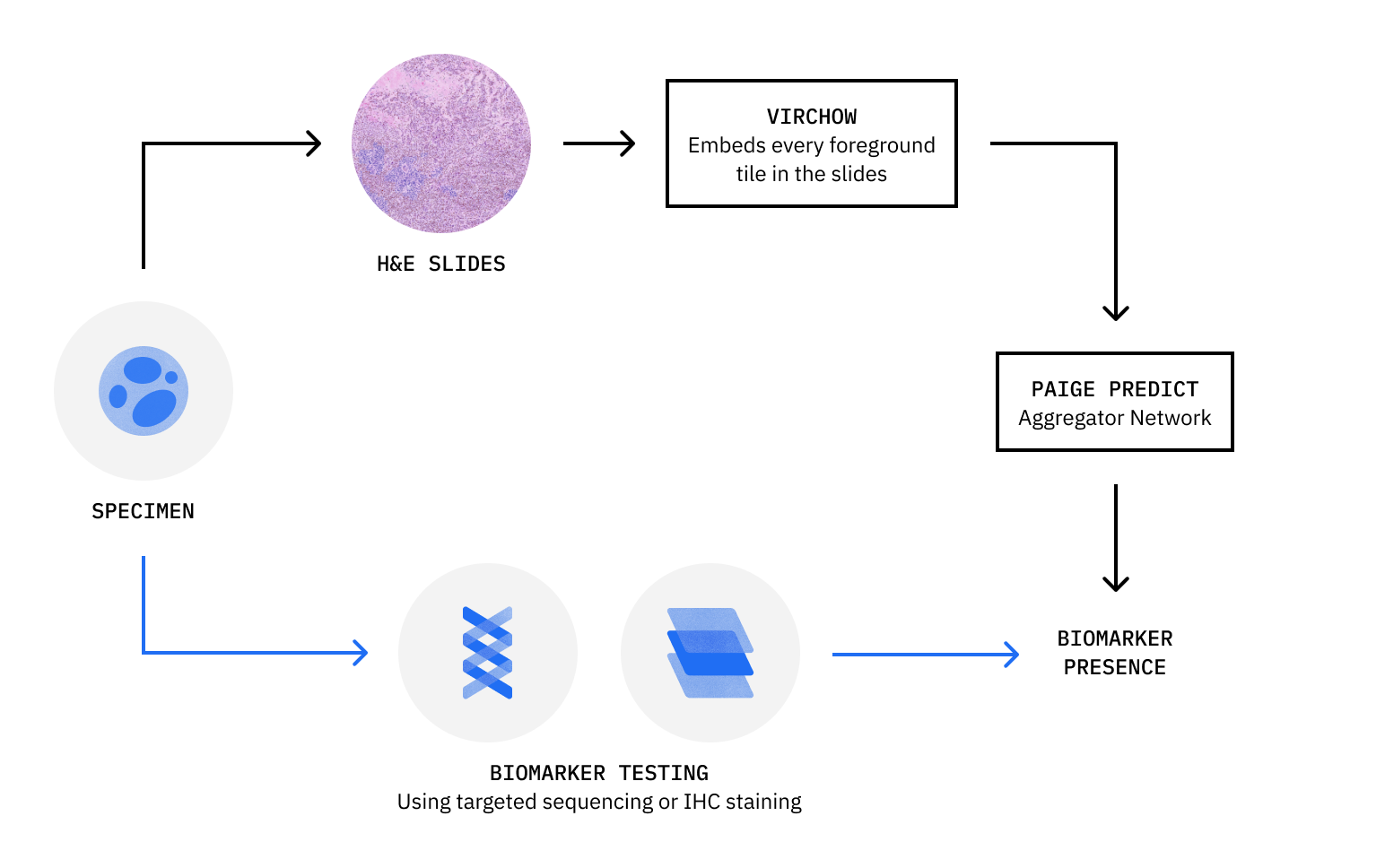
Deep dive: AI in action with Paige Predict
Digital biomarkers have multiple uses. By leveraging Paige Predict to screen for genomic and phenotypic biomarkers, clinicians and researchers have rapid access to valuable information to prioritize testing, pre-screen patients at scale, and enable novel development of AI-enabled diagnostics.
Providing insights when tissue is insufficient for sequencing
In oncology, obtaining sufficient tissue for comprehensive molecular analysis is a significant hurdle. With insufficient tissue, sequencing may deliver a “quantity not sufficient” (QNS) result, lacking actionable insights for patients and providers, resulting in diagnostic delays, and reducing the amount of tissue available for further testing.
xT QNS reports with Paige Predict insights:
Paige Predict is available with approximately 93%6 of our eligible xT QNS reports when tissue-based sequencing is unsuccessful, providing AI insights on the likely status of clinically relevant biomarkers in each patient’s tumor. These insights, which must be confirmed through additional testing, provide crucial information to oncologists as they evaluate further testing decisions, including exploring liquid biopsy panels or targeted, rapid biomarker testing with remaining tissue.
Accelerating biomarker screening to the point of diagnosis
For patients, obtaining molecular testing results with genomic predictions quickly is a critical step in their care journey. Paige Predict is able to analyze the digitized images created as part of the standard diagnostic process, providing AI results in approximately 5 minutes. This speed allows providers to pre-screen samples for genomic information, and identify the confirmatory tests, including NGS, IHC, and targeted gene panels, which are most likely to provide meaningful diagnostic insights to inform a patient’s care pathway.
Enriching cohorts for clinical research at scale
Identifying and enrolling patients for clinical trials is a cost-and-time-intensive hurdle in therapeutic development, often requiring screening hundreds of patients to find just a handful of eligible candidates. Paige Predict can be used in research settings to rapidly pre-screen samples for specific mutations prior to confirmatory testing, supporting efforts to reduce cost and accelerate time to enrollment in clinical studies. Paige Predict enables the rapid, decentralized screening of a significantly larger patient population than currently possible while simultaneously enriching mutation prevalence within that group, helping identify more actionable insights from the same volume of testing.
Building the future of AI-enabled diagnostics
Built with paired pathology and genomic data, Paige Predict can identify both established phenotype-genotype correlations, as well as genomic alterations without known phenotypes. Additionally, Paige Predict reliably identifies activation in groups of genes with shared signaling pathways, providing an opportunity to improve future diagnostic tests by combining both phenotypic and genotypic signatures.
Researchers can use Paige Predict as a technology building block, further extending the model’s capabilities to accelerate the development of multimodal AI applications for novel biomarker identification. As new targeted therapies emerge, this technology serves as a powerful engine for developing AI-enabled companion diagnostics (CDx), in a wide range of use cases, improving response prediction, patient stratification, and informing treatment pathways with speed and precision.
Looking ahead: The future of AI in digital pathology with Tempus
At Tempus, our mission is to build the operating system for cancer care by structuring and analyzing vast amounts of clinical, molecular, and imaging data to make it accessible and useful. We believe that developing novel, multimodal AI applications, validated and made available to clinicians and researchers, is the key to unlocking AI-enabled diagnostics, and realizing the promise of precision medicine, at scale.
The clinical launch of Paige Predict within our lab workflows is a tangible step forward, providing meaningful insights to providers even when tissue is insufficient for sequencing. This is only the beginning. Our vision is to continue expanding the Tempus platform by integrating more diverse data modalities, building more sophisticated AI-enabled research tools, and providing new insights to support precision medicine research. Tempus remains committed to pushing the boundaries of what is possible in precision medicine, building intelligent tools that empower clinicians and researchers to improve the lives of patients everywhere.
Discover the rest of the Paige Predict suite, including Paige Predict for tissue optimization.
For partnerships or to learn more, contact us today.
1. All Paige Predict features other than those available for clinical use with xT CDx are for Research Use Only (RUO) and are not for use in diagnostic procedures. Clinical use of Paige Predict at the Tempus Chicago lab has been validated as a laboratory developed test and is performed for eligible xT QNS cases.
2. Kather JN, Heij LR, Grabsch HI, et al. Pan-cancer image-based detection of clinically actionable genetic alterations. Nature Cancer. 2020;1(8):789–799.
3. Couture HD. Deep Learning-Based Prediction of Molecular Tumor Biomarkers from H&E: A Practical Review. J Pers Med. 2022;12(12):2022.
4. Qu H, Zhou M, Yan Z, et al. Genetic mutation and biological pathway prediction based on whole slide images in breast carcinoma using deep learning. npj Precision Oncology. 2021;5(1):87.
5. Wang YK, Tydlitatova L, Kunz JD, Oakley G, Chow BKK, Godrich RA, et al. Screen Them All: High-Throughput Pan-Cancer Genetic and Phenotypic Biomarker Screening from H&E Whole Slide Images. arXiv. 2024. https://doi.org/10.48550/arXiv.2408.09554
6. Percentage of xT QNS reports that will include Paige Predict results is based on real-world data from 2025 and reflects the cancer cohorts included in the Paige Predict LDT. Samples must be eligible orders and meet specimen requirements.
-
05/05/2026
From recurrence monitoring to treatment response: MRD data insights across the cancer care continuum
This session will explore how minimal residual disease (MRD) data provides critical insights across the cancer care continuum.
Watch replay -
04/02/2026
Unlocking foundation models: Our experience from proof of concept to deployment at scale
An executive discussion on leveraging multi-modal foundation models to accelerate your drug development journey
Watch replay -
03/04/2026
Supporting the financial health of clinical research programs with the Tempus TIME Trial Network
By combining the power of AI-driven technology with a streamlined operational model, Tempus provides the infrastructure and support to enable sites with limited resources to participate in research and expand their capabilities.
Read more


