-
PROVIDERS
New MRD Medicare Coverage for Select Indications*
*When coverage criteria are met. Additional criteria and exceptions for coverage may apply.
-
LIFE SCIENCES
Register now
UPCOMING WEBINAR
Unlocking Foundation Models: Our experience from proof of concept to deploying at scale -
PATIENTS
It's About Time
View the Tempus vision.
- RESOURCES
-
ABOUT US
View Job Postings
We’re looking for people who can change the world.
- INVESTORS
02/24/2026
Q&A: Driving scientific discovery with real-world multimodal data
Paul Fields, PhD, Senior Director of Life Science Strategy at Tempus, discusses how Tempus’ real-world multimodal data and advanced analytics platform can help de-risk and accelerate drug development programs.
Authors
Paul Fields, PhD
Senior Director, Life Science Strategy, Tempus

Senior Director, Life Science Strategy, Tempus

During a recent webinar, Fields highlighted the challenges in bringing precision medicine to the clinic and demonstrated how integrating genomic, transcriptomic, clinical, and imaging data provides a more complete picture of patient care. This approach supports the entire drug development lifecycle, from identifying novel targets and optimizing trial design to helping de-risk late-stage programs.
Q: How does Tempus’ multimodal data support early-stage drug discovery and target identification? |
Fields, PhD: We use our integrated molecular and clinical data to identify potential novel drug targets. In one example in lung adenocarcinoma, we analyzed genomic, transcriptomic, and clinical features to define distinct patient subtypes. These subtypes showed unique molecular profiles and pathogenic mutations, allowing us to dig deeper. By integrating this information with protein-protein interaction networks and patient outcomes, we built a prioritized list of potential targets.
We then validated these targets using our vast library of patient-derived organoids (PDOs). By selecting PDOs that matched the molecular subtypes identified in our real-world data (RWD), we could directly test the impact of knocking out these targets via CRISPR analysis. This process, which connects real-world evidence with preclinical models, allows us to confidently advance the most promising targets into the drug development pipeline. |
Q: Once a lead candidate is identified, how can RWD help optimize clinical trial design? |
Fields, PhD: RWD is invaluable for de-risking a clinical development plan and optimizing Phase 1 and 2 strategies. For example, when developing an antibody-drug conjugate (ADC), you can use our data to understand the expression of your target, like TROP2, across dozens of cancer types. This helps identify indications where the target is most prevalent.
However, it goes beyond simple target expression. You can analyze co-expression with other known biomarkers to inform combination or sequencing strategies. By linking biopsies to a patient’s clinical journey, you can also study expression changes before and after specific therapies. Furthermore, you can analyze signatures related to payload response to get a complete picture of how your drug might perform. This allows you to balance the benefits of an all-comer trial versus a biomarker-selected approach and design inclusion/exclusion criteria that enroll the patients most likely to respond. |
|
“Our primary approach is to work as scientific partners with our collaborators to build a fit-for-purpose dataset that answers a specific research question with integrity.“ – Paul Fields, PhD, Senior Director, Life Science Strategy, Tempus |
Q: Can you provide an example of a novel biomarker developed using Tempus’ data? |
Fields, PhD: A key example is our Immune Profile Score (IPS), a pan-cancer signature designed to predict response to immune checkpoint inhibitors (ICIs). We know that existing biomarkers like PD-L1 and TMB are imperfect, so we sought to improve upon them with a biology-informed approach.
Leveraging the wealth of our data, we integrated information on immune infiltrates, transcriptional signatures, and IO resistance mechanisms. We refined numerous biomarkers down to a final model that uses TMB, the expression of eight specific genes, and three immune signatures to generate a single score from 0 to 100. This score identifies patients with a high probability of responding to ICI therapy. We have since made IPS available to our research partners, allowing them to analyze its interaction with other biomarkers, such as MTAP deletions, to generate new hypotheses for combination therapies. |
Q: How does the Tempus platform enable researchers to analyze this complex data efficiently? |
Fields, PhD: Our Tempus Lens platform provides a secure cloud environment to explore our data. It features a no-code interface that allows researchers to quickly build specific patient cohorts using a wide range of filters, including diagnosis, treatment history, and molecular features.
For more complex analyses, you can move these cohorts into a coding environment. We have built tools that allow investigators to get to insights quickly. For instance, in a live demonstration, we identified a cohort of several hundred urothelial cancer patients treated with a specific ADC/ICI combination. In just a few lines of code, we pulled in their IPS data and overall survival outcomes, demonstrating that high-IPS patients had improved survival. This ability to rapidly move from hypothesis to analysis accelerates the research process significantly. |
Q: How do you ensure the reliability and quality of Tempus’ RWD? |
Fields, PhD: We have put a lot of effort into increasing the quality of our data. Our primary approach is to work as scientific partners with our collaborators to build a fit-for-purpose dataset that answers a specific research question with integrity. This may mean prioritizing cohorts with complete genomic data for discovery research or curating data to be more regulatory-grade for submission discussions.
To build confidence, we also perform internal benchmarking. We compare our data to public sources like SEER and have validated our survival outcomes against those from pivotal clinical trials, showing good concordance. Ultimately, it comes down to scientific trust and providing the evidence that helps our customers build confidence in the data and the insights it generates. |
Q: RWD is often heterogeneous and has missing information. How does Tempus address these challenges? |
Fields, PhD: Missingness is an inherent feature of RWD, and we take a bespoke approach to address it based on the specific research question. In some cases, the best method is to omit records with missing information and focus on a complete dataset. In others, we can prioritize biomarkers that are more complete across the cohort.
For situations with minimal missingness, we can use scientifically sound methods like data imputation. Our data scientists work closely with our customers to determine the best strategy for each project, whether it involves focusing on the most complete data or using an ensemble approach to piece together different sources of information. |
Q: How can the Tempus dataset be used to study treatment resistance? |
Fields, PhD: Our ability to place biopsies within a patient’s full clinical journey is a key advantage for studying both innate and acquired resistance. To study innate resistance, we can compare pre-treatment biopsies from patients who have a long-term response to a therapy versus those who experience immediate progressive disease.
To understand acquired resistance, we can analyze biopsies collected after a patient has relapsed. This allows us to identify molecular changes, such as the emergence of an ESR1 mutation, that may be driving resistance. This information is critical for positioning a drug, helping decide whether it should be used in combination with a prior line of therapy or as a monotherapy after treatment failure. |
Note: Content edited for clarity. Please note that the content in this document has been revised for clarity and conciseness. Some language and formatting may have been adjusted to enhance readability while preserving the original meaning and intent of the discussion.
|
To gain comprehensive insights into Tempus’ role in helping power new hypotheses and accelerate discovery, and to see Tempus Lens in action, we invite you to watch the webinar recording. For in-depth demonstrations of our AI-enabled applications, contact us.
|
-
03/24/2026
Translating data into an actionable R&D strategy
Hear from industry leaders about how a data-driven understanding of disease biology informs critical decisions and accelerates therapeutic development. Secure your recording now.
Watch replay -
03/17/2026
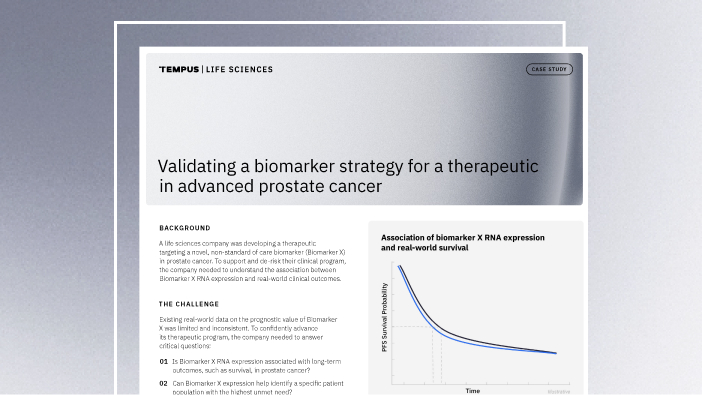
Validating a biomarker strategy for a therapeutic in advanced prostate cancer
Download the case study to learn how Tempus helped a life sciences partner use real-world data to clarify a biomarker’s value, define a high-risk patient population, and advance their therapeutic program with greater confidence.
Read more -
03/09/2026
From model to medicine: Closing the preclinical to clinical gap with an integrated, data-driven engine
Richard Klinghoffer, SVP and Head of Systems Biology at Tempus, discusses how preclinical models can be disconnected from real patient biology, contributing to challenges in oncology drug development.
Read more